Full article: RRM3 regulates epigenetic conversions in Saccharomyces cerevisiae in conjunction with Chromatin Assembly Factor I

High-Affinity Binding of Chp1 Chromodomain to K9 Methylated Histone H3 Is Required to Establish Centromeric Heterochromatin: Molecular Cell

Central role of Drosophila SU(VAR)3–9 in histone H3‐K9 methylation and heterochromatic gene silencing | The EMBO Journal
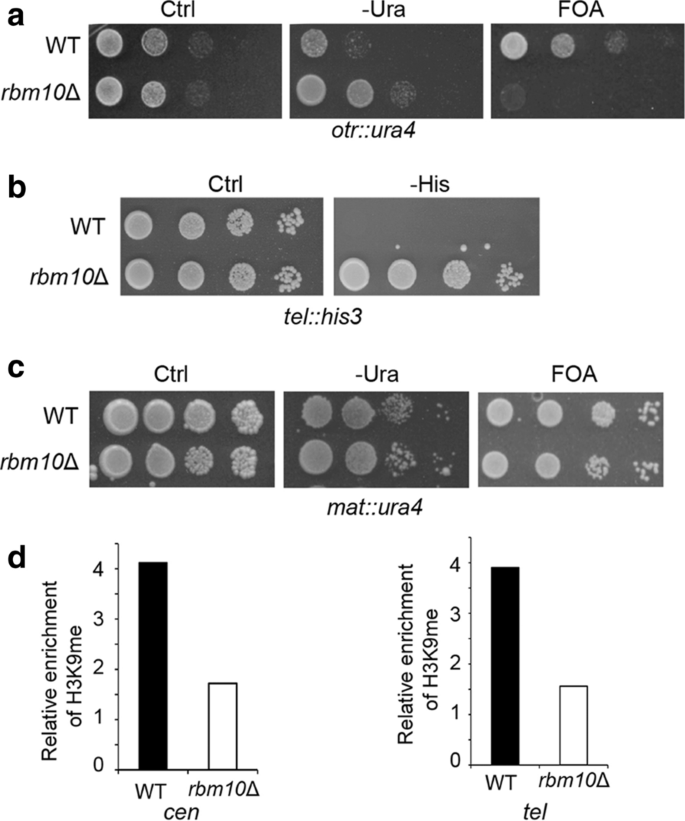
Rbm10 facilitates heterochromatin assembly via the Clr6 HDAC complex | Epigenetics & Chromatin | Full Text

Full article: RRM3 regulates epigenetic conversions in Saccharomyces cerevisiae in conjunction with Chromatin Assembly Factor I

A Heterochromatin Barrier Partitions the Fission Yeast Centromere into Discrete Chromatin Domains - ScienceDirect
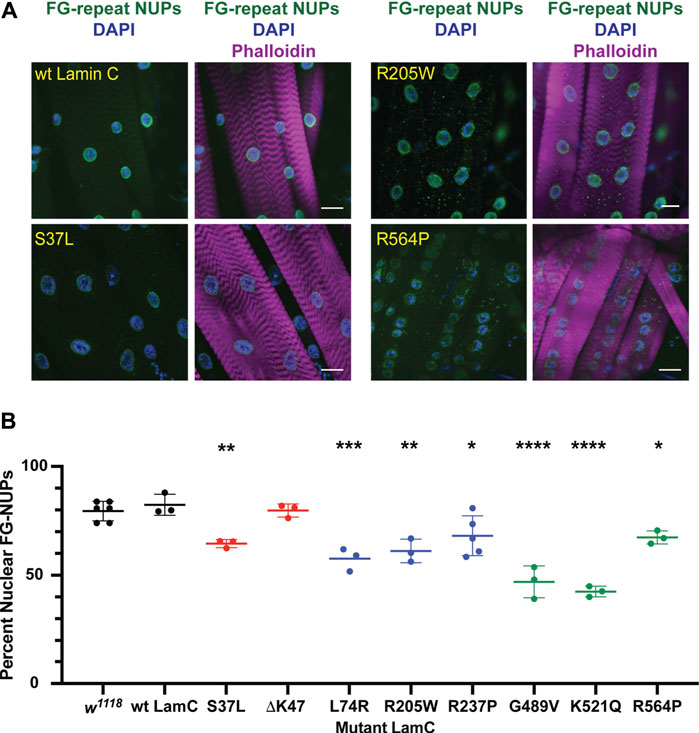
Frontiers | Effects of mutant lamins on nucleo-cytoskeletal coupling in Drosophila models of LMNA muscular dystrophy

siRNA-Mediated Heterochromatin Establishment Requires HP1 and Is Associated with Antisense Transcription - ScienceDirect
Fission yeast CENP-B homologs nucleate centromeric heterochromatin by promoting heterochromatin-specific histone tail modificati

Central role of Drosophila SU(VAR)3-9 in histone H3-K9 methylation and heterochromatic gene silencing. - Abstract - Europe PMC

Silencing in Yeast rDNA Chromatin: Reciprocal Relationship in Gene Expression between RNA Polymerase I and II - ScienceDirect

Moške Limited Edition Gledati Neodvisno Watchmaker Dela S Polnim Automatic Mehanski Nerjavečega Jekla Nepremočljiva Retro Punk Watch nakup ~ Ure < www.ful-povezani.si
The SCFDia2 Ubiquitin E3 Ligase Ubiquitylates Sir4 and Functions in Transcriptional Silencing | PLOS Genetics
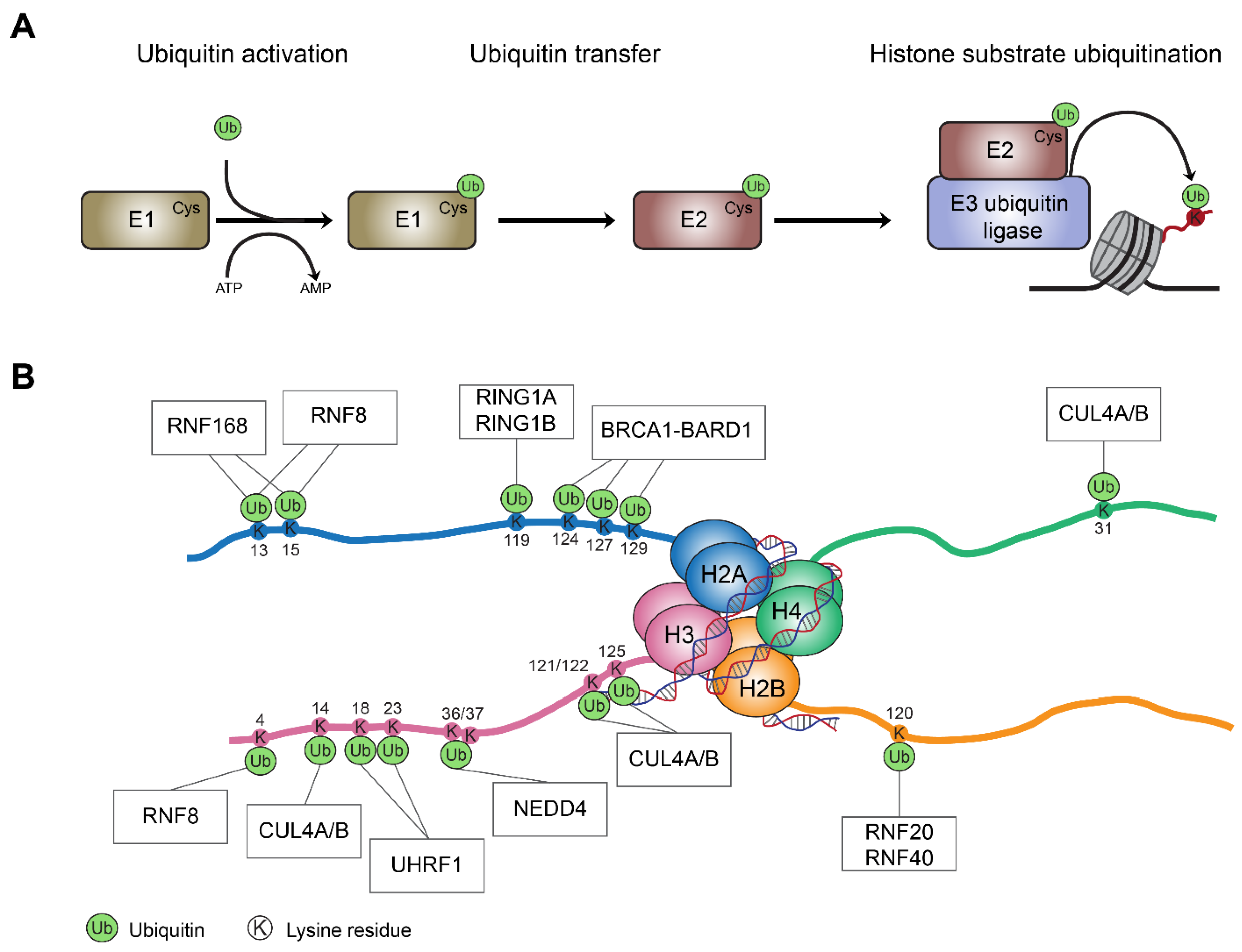
Cells | Free Full-Text | Histone Mono-Ubiquitination in Transcriptional Regulation and Its Mark on Life: Emerging Roles in Tissue Development and Disease

A Heterochromatin Barrier Partitions the Fission Yeast Centromere into Discrete Chromatin Domains - ScienceDirect